trainer.lib package¶
Subpackages¶
Submodules¶
trainer.lib.DslSemantics module¶
-
class
trainer.lib.DslSemantics.DslSemantics(max_resources=10000)¶ Bases:
abc.ABCDefines execution context and utility functionality for running python statements from strings.
Child classes define methods that can be called inside of the DSL constructs.
-
bind_object(o: Any)¶
-
compile_prog(prog: str)¶
-
execute_program(state: Dict) → Generator¶
-
static
gen_wrapper(f: Callable) → _GenericAlias¶ For example, converts a function f: bool -> int to a generator f: Generator[bool] -> Generator[int] :param f: A callable :return: A generator with the semantics of f
-
static
generate_enum(e: enum.EnumMeta) → Generator¶
-
static
is_callable(f: Any) → bool¶
-
trainer.lib.config module¶
The config of trainer is stored in the user directory in a folder ‘.trainer’.
The database connection string should be in the file configfile accessible by the key ‘db_con’: postgresql+psycopg2://postgres:password@127.0.0.1:5432/db_name
-
trainer.lib.config.load_config_json()¶
-
trainer.lib.config.save_config_json(obj=None)¶
trainer.lib.data_model module¶
Data Model¶
The data model aims to simplify machine learning on complex data structures. For example, classifying a subject (medical patient) by both its gender and between 1 and 4 ultrasound videos.
A dataset contains:
Subjects (Which are the training examples)
Model Weights
Json config files - Class definitions - Segmentation mask definitions
A Subject contains:
Images & masks
Classes
Json config files
Multiple different types of binaries are supported.
Image stacks are used for images, videos and 3D images. Shape of an image stack: [#frames, width, height, #channels]
Segmentation Masks (‘img_mask’) are used to store every annotated structure for one frame of an imagestack. Shape of a mask: [width, height, #structures]
Miscellaneous objects are general pickled objects.
-
class
trainer.lib.data_model.ClassDefinition(**kwargs)¶ Bases:
sqlalchemy.ext.declarative.api.Base-
classmethod
build_new(name: str, cls_type: trainer.lib.data_model.ClassType, values: List[str])¶
-
cls_type¶
-
id¶
-
name¶
-
values¶
-
classmethod
-
class
trainer.lib.data_model.ClassType(value)¶ Bases:
enum.EnumAn enumeration.
-
Binary= 'binary'¶
-
Nominal= 'nominal'¶
-
Ordinal= 'ordinal'¶
-
-
class
trainer.lib.data_model.Classifiable¶ Bases:
object-
classes= Column(None, JSONB(astext_type=Text()), table=None)¶
-
get_class(class_name: str)¶
-
classmethod
query_all_with_class(session: sqlalchemy.orm.session.Session, class_name: str)¶
-
remove_class(class_name: str)¶
-
set_class(class_name: str, class_val: str)¶
-
-
class
trainer.lib.data_model.Dataset(**kwargs)¶ Bases:
sqlalchemy.ext.declarative.api.BaseA dataset is a collection of splits.
-
add_split(split_name: str) → trainer.lib.data_model.Split¶
-
classmethod
build_new(name: str)¶
-
get_split_by_name(split_name: str)¶
-
get_summary() → str¶
-
id¶
-
name¶
-
splits: List[Split]¶
-
-
class
trainer.lib.data_model.ImStack¶ Bases:
trainer.lib.data_model.Classifiable,trainer.lib.data_model.NumpyBinary,sqlalchemy.ext.declarative.api.Base-
add_ss_mask(gt_arr: numpy.ndarray, sem_seg_tpl: trainer.lib.data_model.SemSegTpl, for_frame=0, ignore_shape_mismatch=False)¶
-
binary¶
-
classmethod
build_new(src_im: numpy.ndarray, extra_info: Optional[Dict] = None)¶ Only adds images, not volumes or videos! Unless it is already in shape (frames, width, height, channels). Multi-channel images are assumed to be channels last. Grayscale images are assumed to be of shape (width, height).
The array is saved using type np.uint8 and is expected to have intensities in the range of [0, 255]
- Parameters
src_im – Numpy Array. Can be of shape (W, H), (W, H, #C) or (#F, W, H, #C)
extra_info – Extra info for a human. Must contain only standard types to be json serializable
-
classes¶
-
dtype¶
-
extra_info¶
-
file_path¶
-
id¶
-
sbjt_id¶
-
semseg_masks: List[SemSegMask]¶
-
shape¶
-
stored_in_db¶
-
-
class
trainer.lib.data_model.MaskType(value)¶ Bases:
enum.EnumPossible types that a mask can have.
blob: straightforward region. Is used for most segmentation tasks
A point is usually segmented as a small circle and then postprocessed to be the center of that circle
A line is usually segmented as a sausage and then postprocessed to a single response-line
-
Blob= 'blob'¶
-
Line= 'line'¶
-
Point= 'point'¶
-
Unknown= 'unknown'¶
-
class
trainer.lib.data_model.NumpyBinary¶ Bases:
object-
binary= Column(None, LargeBinary(), table=None)¶
-
dtype= Column(None, String(), table=None)¶
-
file_path= Column(None, String(), table=None)¶
-
get_bin_disk_path()¶
-
init_on_load()¶ Does the job of the constructor in case of an object which is loaded from the database. See https://docs.sqlalchemy.org/en/13/orm/constructors.html for details.
-
set_array(arr: numpy.ndarray) → None¶
-
shape= Column(None, String(), table=None)¶
-
stored_in_db= Column(None, Boolean(), table=None)¶
-
values() → numpy.ndarray¶
-
-
class
trainer.lib.data_model.SemSegClass(**kwargs)¶ Bases:
sqlalchemy.ext.declarative.api.Base-
classmethod
build_new(name: str, ss_type: trainer.lib.data_model.MaskType)¶
-
id¶
-
name¶
-
ss_type¶
-
tpl_id¶
-
classmethod
-
class
trainer.lib.data_model.SemSegMask¶ Bases:
trainer.lib.data_model.Classifiable,trainer.lib.data_model.NumpyBinary,sqlalchemy.ext.declarative.api.Base-
binary¶
-
classes¶
-
dtype¶
-
file_path¶
-
for_frame¶
-
id¶
-
im_stack_id¶
-
mtype¶
-
shape¶
-
stored_in_db¶
-
tpl¶
-
tpl_id¶
-
-
class
trainer.lib.data_model.SemSegTpl(**kwargs)¶ Bases:
sqlalchemy.ext.declarative.api.Base-
classmethod
build_new(tpl_name: str, seg_types: Dict[str, trainer.lib.data_model.MaskType])¶
-
id¶
-
name¶
-
ss_classes: List[SemSegClass]¶
-
classmethod
-
class
trainer.lib.data_model.Split(**kwargs)¶ Bases:
sqlalchemy.ext.declarative.api.Base-
dataset_id¶
-
id¶
-
name¶
-
sbjts: List[Subject]¶
-
-
class
trainer.lib.data_model.Subject(**kwargs)¶ Bases:
trainer.lib.data_model.Classifiable,sqlalchemy.ext.declarative.api.BaseIn a medical context a subject is concerned with the data of one patient. For example, a patient has classes (disease_1, …), imaging (US video, CT volumetric data, x-ray image, …), text (symptom description, history) and structured data (date of birth, nationality…).
The extra_info attribute can be used freely for a json dict.
In future releases a complete changelog will be saved in a format suitable for process mining.
-
classmethod
build_new(name: str, extra_info: Optional[Dict] = None)¶
-
classes¶
-
extra_info¶
-
id¶
-
ims: List[ImStack]¶
-
name¶
-
classmethod
-
trainer.lib.data_model.reset_data_model()¶
trainer.lib.gen_utils module¶
-
class
trainer.lib.gen_utils.GenCacher(*args, **kwds)¶ Bases:
typing.GenericWrapper around a generator that stores the already yielded values and therefore allows indexing.
-
fill_cache(idx: int)¶
-
get_cache_len() → int¶
-
is_exhausted() → bool¶
-
-
trainer.lib.gen_utils.product(gens: List[Generator]) → Generator¶ Utility to compute the cartesian product between an arbitrary number of generators. Developed to handle the case of a possible mix of finite and infinite generators. The built-in itertools.product can only compute the cartesian product between finite generators.
The exploration strategy can be visualized using the following code block:
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27
import matplotlib.pyplot as plt import trainer.demo_data as dd import trainer.lib as lib fig = plt.figure() ax = fig.add_subplot(111, projection='3d') ax.set_xlim3d([0.0, 10.0]) ax.set_xlabel('X') ax.set_ylim3d([0.0, 10.0]) ax.set_ylabel('Y') ax.set_zlim3d([0.0, 10.0]) ax.set_zlabel('Z') xs, ys, zs = [], [], [] gens = [ dd.finite_test_gen(start=0, end=3), dd.infinite_test_gen(first=0), dd.finite_test_gen(start=0, end=3) ] for c in lib.product(gens): xs.append(c[0]) ys.append(c[1]) zs.append(c[2]) ax.plot(xs=xs[-2:], ys=ys[-2:], zs=zs[-2:]) fig.show() plt.pause(0.01)
The result looks as following:
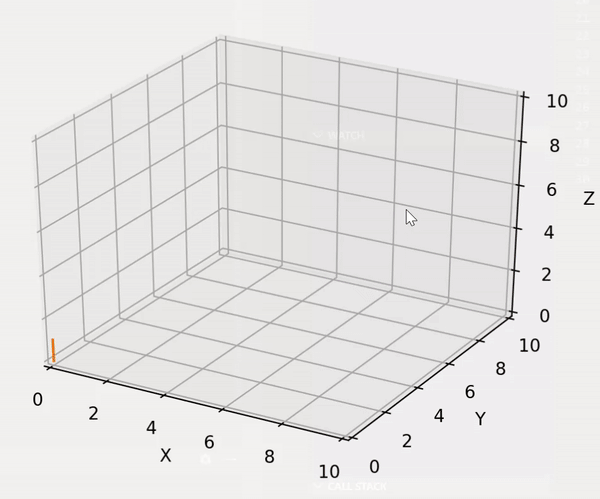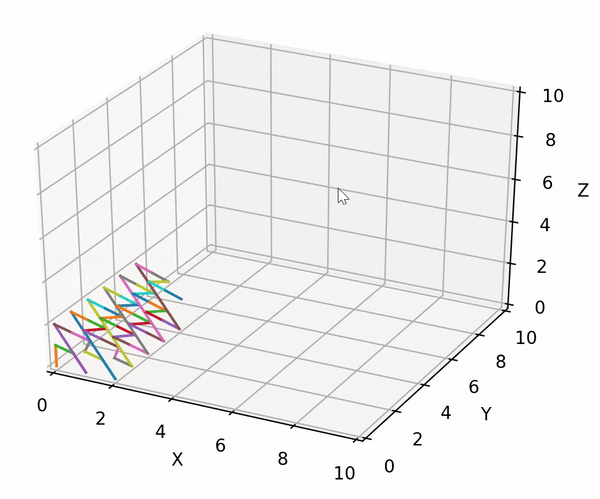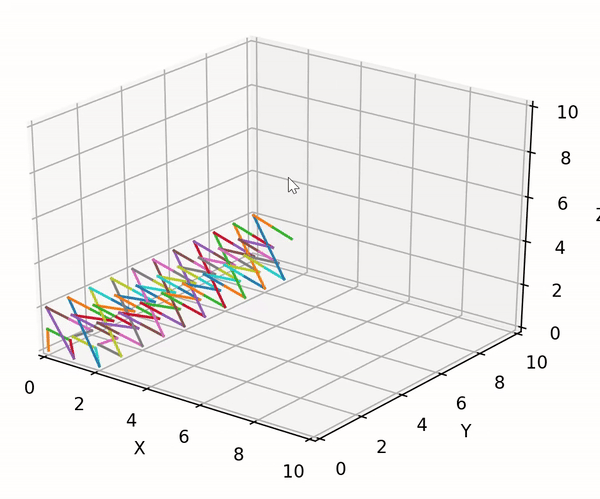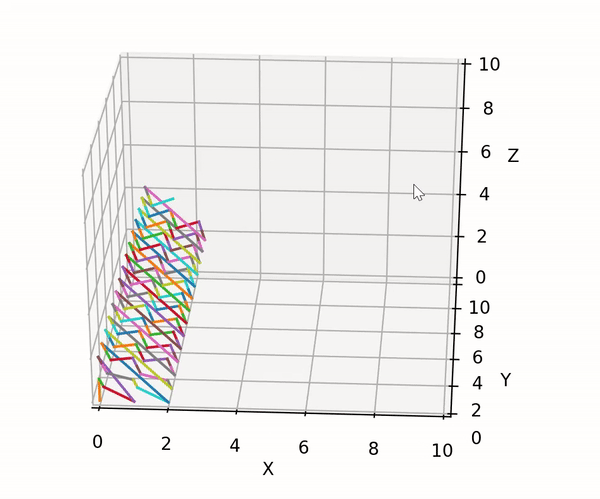
- Parameters
gens – Between 1 and N generators
- Returns
One generator that returns all N-tuples, built from the input generators
-
trainer.lib.gen_utils.sample_randomly(gens: Union[List[Generator], List[Iterator]], probas: List[float], use_softmax=False)¶ Draw from one generator in a list according to uniformly distributed probabilities.
- Parameters
gens – A list of generators
probas – List of generator probabilities, must correspond to the list of generators
use_softmax – Use softmax to press priorities to one
- Returns
Randomly drawn value from one of the generators
-
trainer.lib.gen_utils.summations(sum_to: int, ls: List[int]) → Generator[Tuple, None, None]¶
trainer.lib.grammar module¶
-
class
trainer.lib.grammar.Grammar(*args, **kwds)¶ Bases:
typing.Generic-
append_semantics(f: Callable, prio: float)¶
-
get_rule(nts: NTS) → List[Tuple[List[Union[TS, NTS]], float]]¶
-
read_program(start_symbol: NTS) → Generator[Union[List[TS], None]]¶
-
sample_prog_strings(sym: NTS)¶
-
-
trainer.lib.grammar.analyse_function_type(f: Callable) → Tuple[List[type], type]¶
trainer.lib.import_utils module¶
Collection of utility function which can be used to add content from other file-formats and sources to the convenient trainer dataset-format.
-
trainer.lib.import_utils.add_image_folder(split: trainer.lib.data_model.Split, folder_path: str, progress=True, sess=<sqlalchemy.orm.session.Session object>) → None¶ Iterates through a folder and adds its contents to a split.
If a file is found, a new subject is created with only that file. If a directory is found, a new subject is created with all files that live within that directory. If a dicom file is found, the image is appended to the subject with that patient_id
Supported file formats: - DICOM (no extension or .dcm) - Standard image files - B8 files (.b8)
- Parameters
split – The dataset split this data is appended to. The split knows its dataset.
folder_path – Top level folder path
progress – If true, displays a progress bar
sess – database session, defaults to a new session
-
trainer.lib.import_utils.add_imagestack(s: trainer.lib.data_model.Subject, file_path: str, binary_id='') → None¶ Takes an image path and tries to deduce the type of image from the path ending. No path ending is assumed to be a DICOM file (not a DICOM folder)
-
trainer.lib.import_utils.add_import_folder(split: trainer.lib.data_model.Split, folder_path: str, semsegtpl: trainer.lib.data_model.SemSegTpl)¶
-
trainer.lib.import_utils.export_to_folder(split: trainer.lib.data_model.Split, folder_path: str)¶
-
trainer.lib.import_utils.import_dicom(dicom_path: str)¶
-
trainer.lib.import_utils.import_subject(split: trainer.lib.data_model.Split, subject_path: str, semsegtpl: trainer.lib.data_model.SemSegTpl)¶ Imports a subject from the format created when exporting datasets
trainer.lib.logging module¶
-
class
trainer.lib.logging.Experiment(**kwargs)¶ Bases:
sqlalchemy.ext.declarative.api.BaseAllows to track solutions over time. Uses the database for different types of logs.
-
add_result(result_name: str, flag='success', sess=None, auto_commit=True) → None¶ Add a new result to the current run.
- Parameters
result_name – Value of the result.
flag – Flag of the result. For example ‘success’ or ‘fail’. It is case-sensitive.
sess –
auto_commit –
-
classmethod
build_new(experiment_name: str, sess=None)¶
-
experiment_name¶
-
static
get_all_with_flag(sess: sqlalchemy.orm.session.Session, exp_name: str, flag='success') → List[str]¶ Computes all results with a certain flag from the history i.g. all runs.
-
get_results(flag='success') → List[str]¶
-
id¶
-
is_in(result_name: str, flag='success') → bool¶ Returns if the result was already added to the current experiment.
-
results: List[ExperimentResult]¶
-
start_date¶
-
-
class
trainer.lib.logging.ExperimentResult(**kwargs)¶ Bases:
sqlalchemy.ext.declarative.api.BaseThe semantics of one instance of this class might be: data point #223 was correctly classified.
-
classmethod
build_new(name: str, flag: str)¶
-
exp_id¶
-
flag¶
-
id¶
-
name¶
-
classmethod
-
class
trainer.lib.logging.LogWriter(log_dir: str = './logs', id_hint='log')¶ Bases:
object-
debug_var(o: Any) → None¶ Allows to inspect an arbitrary python object on disk.
For visualizing an array with a description debug a (np.ndarray, str) tuple. For visualizing multiple arrays with one description each debug a (np.ndarray, str) list.
- Parameters
o – Any variable
-
get_absolute_run_folder() → str¶
-
get_parent_log_folder() → str¶
-
trainer.lib.misc module¶
-
trainer.lib.misc.create_identifier(hint: str = '') → str¶ Can be used to create unique names for files by exploiting the uniqueness of the current date. Be aware that if two identifiers are created during the same second they are equal! Follows the form YYYY_MM_DD__hh_mm_ss.
- Returns
YYYY_MM_DD__hh_mm_ss_{hint}
-
trainer.lib.misc.delete_dir(dir_path: str, blocking=True, verbose=True)¶
-
trainer.lib.misc.download_and_extract(online_url: str, parent_dir='./', dir_name: str = None) → str¶ Can be used to download and extract a .zip dataset file hosted online. Assumes the zip to be one directory. :param online_url: The url that points directly to a .zip file containing folders with the dataset files :param parent_dir: The directory that is used to store the temporary zip and the final extracted folder :param dir_name: if provided, the function checks if the directory already exists and does not download it again :return: The absolute local path to the directory
-
trainer.lib.misc.get_img_from_fig(fig: matplotlib.pyplot.figure, dpi=180) → numpy.ndarray¶ Converts a matplotlib figure into a numpy array.
- Parameters
fig – A matplotlib figure
dpi – image quality, higher is better and takes longer
- Returns
np.ndarray with the image content
-
trainer.lib.misc.load_b8(file_path: str) → numpy.ndarray¶ Loads b8 file used by some ultrasound machines.
- Param
file_path: The direct path to the b8 file
- Returns
Numpy array with the image data
-
trainer.lib.misc.load_grayscale_from_disk(path: str) → numpy.ndarray¶
-
trainer.lib.misc.make_converter_dict_for_enum(e)¶
-
trainer.lib.misc.pick_from_list(ls: List[T], title='Title', rows=- 1, columns=- 1) → T¶ Spawns a small gui which allows the user to select from a list.
Be aware that this works only if a $Display is set
-
trainer.lib.misc.slugify(value)¶ Normalizes string, converts to lowercase, removes non-alpha characters, and converts spaces to hyphens.
-
trainer.lib.misc.standalone_foldergrab(folder_not_file: bool = False, optional_inputs: List[Tuple[str, str]] = None, optional_choices: List[Tuple[str, str, List[str]]] = None, title='Select a folder') → Tuple[str, Dict[str, str]]¶ Uses a simple blocking GUI for prompting the user for a file or folder path. Optionally allows to prompt for additional text inputs as well.
- Parameters
folder_not_file – if True asks for a folder, if False asks for a file
optional_inputs – A list of (description, key) pairs
optional_choices – A list of (description, key, list of options) pairs
title –
- Returns
Tuple with path as first entry and a dictionary with the optional text inputs as second entry
trainer.lib.syn module¶
Given rules of a grammar the syn package searches for syntactic correct solutions for example input/output pairs.
-
class
trainer.lib.syn.NonTerminalSym(rule: trainer.lib.syn.SubstitutionRule)¶ Bases:
object
-
class
trainer.lib.syn.State(pair: Pair)¶ Bases:
object-
is_final() → bool¶ Can be used to determine for one
:return True if the state is a valid solution to the output
-
visualize(ax1, ax2)¶
-
-
class
trainer.lib.syn.SubstitutionRule(left_side, right_side)¶ Bases:
object
-
class
trainer.lib.syn.TerminalSym¶ Bases:
object-
apply_on_state(s: trainer.lib.syn.State)¶
-
semantics()¶
-